The HSLS Molecular Biology Information Service recently licensed two new bioinformatics resources from CLC bio. CLC Main Workbench supports researcher’s daily bioinformatics needs and CLC Genomics Workbench handles sequencing data from high-throughput sequencing systems.
CLC Main Workbench is an integrated software package that enables users to perform advanced DNA, RNA, and protein sequence analyses, combined with gene expression analysis, seamless data management, and user-friendly graphical viewing and output options. Noteworthy features include but are not limited to:
- Expression analysis including digital gene expression: support for both microarray and sequencing-based expression data, clustering algorithms, tools for Gene Set Enrichment Analysis
- DNA sequence analysis: primer design, in silico PCR, cloning, SNP annotation, restriction enzyme analysis and management
- RNA structure analysis: secondary structure prediction, symbolic representations
Protein sequence analysis: antigenicity, PFAM domain search, transmembrane helix prediction, proteolytic cleavage detection - Pattern search: motif searches for basic patterns, using regular expressions, and/or with ProSite patterns
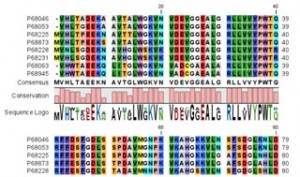
- Database searches: GenBank Entrez, BLAST on local databases, PubMed
Project and data management: detailed history log, full integration of data input, data management, calculation results, and data export - Other bioinformatics features: multiple alignment of DNA, RNA, and proteins, sequence editor, local complexity region analyses
CLC Genomics Workbench is a cross-platform desktop application that incorporates cutting-edge technology and algorithms for analyzing and visualizing next generation sequencing data. Some of its key applications are:
 Genomics: whole genome re-sequencing and targeted re-sequencing of genomes of any size and type, de novo assembly of an unlimited number of reads, SNP and DIP detection, identification of genomic rearrangements
Genomics: whole genome re-sequencing and targeted re-sequencing of genomes of any size and type, de novo assembly of an unlimited number of reads, SNP and DIP detection, identification of genomic rearrangements- Transcriptomics: digital gene expression based on RNA-Seq, including a wide range of downstream gene expression analyses, novel transcript/exon discovery, interactive view of assemblies and derived gene expression data
- Epigenomics: ChIP-seq analysis, peak finding and refinement, false discovery rate and background distribution tables and graphs
To access CLC Main Workbench or CLC Genomics Workbench, visit Licensed Tools on the HSLS Molecular Biology portal.
For more information about CLC bio resources, call Ansuman Chattopadhyay at 412-648-1297, Carrie Iwema at 412-383-6887, or contact Ask A MolBio Librarian.
Parts of this article were reprinted from CLC bio.
~ Carrie Iwema