The HSLS Molecular Biology Information Service (HSLS-MBIS), in collaboration with the Center for Research Computing, is excited to announce the availability of a limited number of licenses for a long-awaited and powerful tool for processing and analyzing single cell RNA-Seq (scRNA-Seq) data—Partek® Flow®.
What is scRNA-Seq? It’s a revolutionary next-generation sequencing technology that focuses on the characterization of individual cells, as opposed to bulk populations of cells. This allows biomedical researchers to make previously challenging discoveries such as the elucidation of gene regulatory networks, the reconstruction of cell hierarchies, and the identification of rare cell types.
Partek® Flow® is a commercially-licensed tool that helps researchers to:
- Process data generated via single cell technology: Partek® Flow® analysis begins with a FASTQ file or a count matrix from 10x Genomics or Drop-Seq. Preset filters recognize and exclude unique molecular identifiers and barcodes from alignment against the reference genome. Additional filters remove low-quality cells and uninformative genes, identify outliers using interactive QA/QC plots, and help identify genes of interest for downstream analysis.
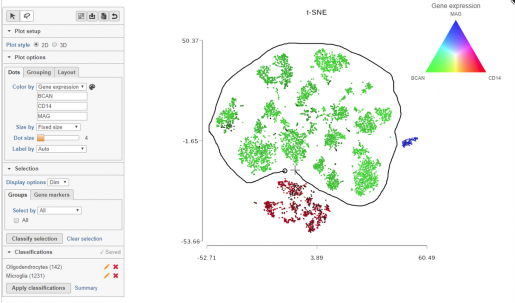
- Discover cell types within single cell data: Partek® Flow® uses an interactive dimensional reduction algorithm, t-distributed stochastic neighbor embedding (t-SNE), to discover cell types within single cell data via the grouping of similar cells as clusters on a scatter plot. Statistical tests within the software enable the discovery and prevalence of differentially expressed genes and pathways between cell types and/or experimental conditions.
- Visualize single cell results: Partek® Flow® provides customizable visualizations. The t-SNE results are viewable in 2D or 3D to show clustering and/or gene expression patterns in different cell types. Hierarchical clustering heat maps and violin plots can be customized as publication-quality SVG files.
Partek® Flow® provides user support in the form of webinars and tutorials. Anyone with a University of Pittsburgh e-mail address can register for access to Partek® Flow® via the HSLS license. HSLS-MBIS will be offering an introductory lecture on single cell RNA-Seq analysis using Partek® Flow® on March 6; you can register for the class here. Contact the HSLS Molecular Biology Information Service with any questions.
~Carrie Iwema
*Parts of this article were derived from the Partek® website.